Mitigating Risk and Ensuring Consistent Supply Chain Through Internal
SARS-CoV-2 Testing with RT-LAMP
by Nathan A. Tanner, Ph.D., Zhiru Li, Ph.D., Yinhua Zhang, Ph.D., Bradley Langhorst, Ph.D., William E. Jack, Ph.D.,
and Clotilde K. S. Carlow, Ph.D., New England Biolabs, Inc.
The COVID-19 pandemic posed a significant threat to the functioning of laboratories and workplaces worldwide, including here at New England Biolabs (NEB®) and our biotechnology operations in Massachusetts. As the pandemic disruptions began to take hold, it was essential to maintain our ability to produce critical materials for molecular diagnostic tests and mRNA vaccine development. Although the traditional clinical diagnostic supply chain quickly became stretched, with nasopharyngeal (NP) swabs and kits for automated RNA extraction in high demand, we were able to leverage unique resources at NEB, including >35 years of experience in infectious disease diagnostics and industry-leading expertise in simple molecular assays. As described here, we established a CLIA-certified onsite testing program utilizing saliva and colorimetric loopmediated isothermal amplification (LAMP), and to date we have processed over 35,000 SARS-CoV-2 tests for our employees. This testing program has been an invaluable tool in keeping our workplaces and staff safe, ensuring our ability to support the global biotechnology community that relies on our reagents and products.
Our first COVID screening program utilized an outside testing provider, Color Health, who were the first to obtain an Emergency Use Authorization from the FDA for an RT-LAMP test (1). This test utilized NEB reagents and was extremely easy to use. However, it required shipping of nasal swabs to a testing lab, which occasionally resulted in delays or failures as materials were held longer than acceptable times due to weather and other unforseen circumstances. Our research groups were already evaluating the compatibility of our colorimetric LAMP technology with saliva, so we increased focus on this work through an Institutional Review Board (IRB) program with volunteer employees and family members. Sample collection and treatment options were tested and refined under the IRB and the diverse sample pool it allowed us to access. However, to enable release of test results to all individual employees and contractors, we went through the process of CLIA certification for our laboratory and test, adopting all training, operation, and documentation requirements for CLIA operation.
Why use RT-LAMP?
NEB has long been a leader in enzymes for molecular diagnostics, with isothermal amplification and LAMP of particular focus. Since its development in the early 2000s (2), LAMP has shown shown the potential to broaden access to diagnostic technology, bringing these methods outside the clinical laboratory to field and point-of-care settings. NEB has facilitated this shift by developing the first WarmStart® enzymes and colorimetric LAMP master mixes, and also by demonstrating the utility of these tools in our longstanding parasitology and neglected tropical disease research (3-5). LAMP also displays a remarkable tolerance for direct sample inputs without requiring nucleic acid purification, a critical benefit for testing outside the traditional clinical diagnostic infrastructure. With our LAMP expertise and its ability to enable rapid, simple diagnostic tests, we naturally thought of colorimetric LAMP as a new approach to SARS-CoV-2 testing. In February 2020, we quickly worked with a partner lab in Wuhan, China to demonstrate the ability of colorimetric LAMP to detect SARS-CoV-2 directly from nasopharyngeal swab samples (6). While this was a very limited and preliminary study, it demonstrated the strong potential for simplified testing. Immediately following this work, we continued optimizing the conditions and assay (7) to create a reliable colorimetric LAMP testing kit for SARSCoV-2 (SARS-CoV-2 Rapid Colorimetric LAMP Assay Kit, NEB #E2019), and with this product in hand, looked to utilize it for in-house testing.
Why Saliva?
We chose to focus on saliva for our test to avoid the NP swab supply bottleneck, and also to avoid a requirement for nucleic acid extraction from the transport media sample, making the workflow as rapid as possible. Saliva collection is simple, with no swabbing required, and is preferred by our employees. Our colleagues at Yale University had pioneered the SalivaDirect workflow (8), and similar extraction-free approaches were beginning to be demonstrated for LAMP (9-10). While dedicated saliva collection devices exist, we saw no need to take those robust, transport-oriented solutions, and instead utilized a collection kit from laboratory supplies present in abundance in our facility: a 1.5 ml screw-cap tube and a 1 ml unfiltered micropipette tip for accurately and cleanly directing saliva into the tube. Employees scan QR code labels affixed to the tubes with their smartphone, enabling the tracking of samples for each registered user, and deposit their individual 1.5 ml tube into 96-well tube racks for collection.
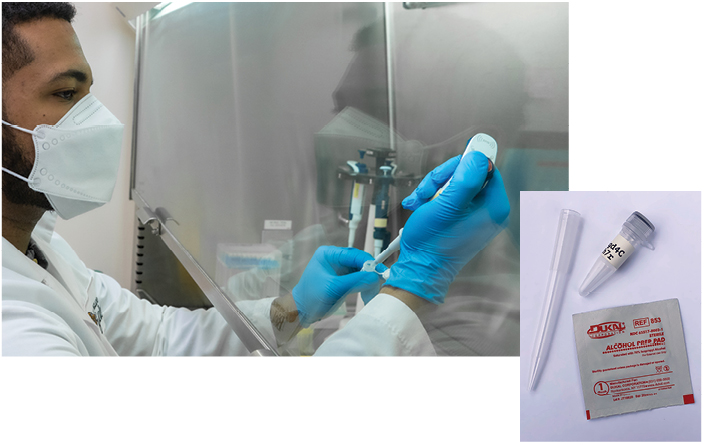

Testing workflow outline
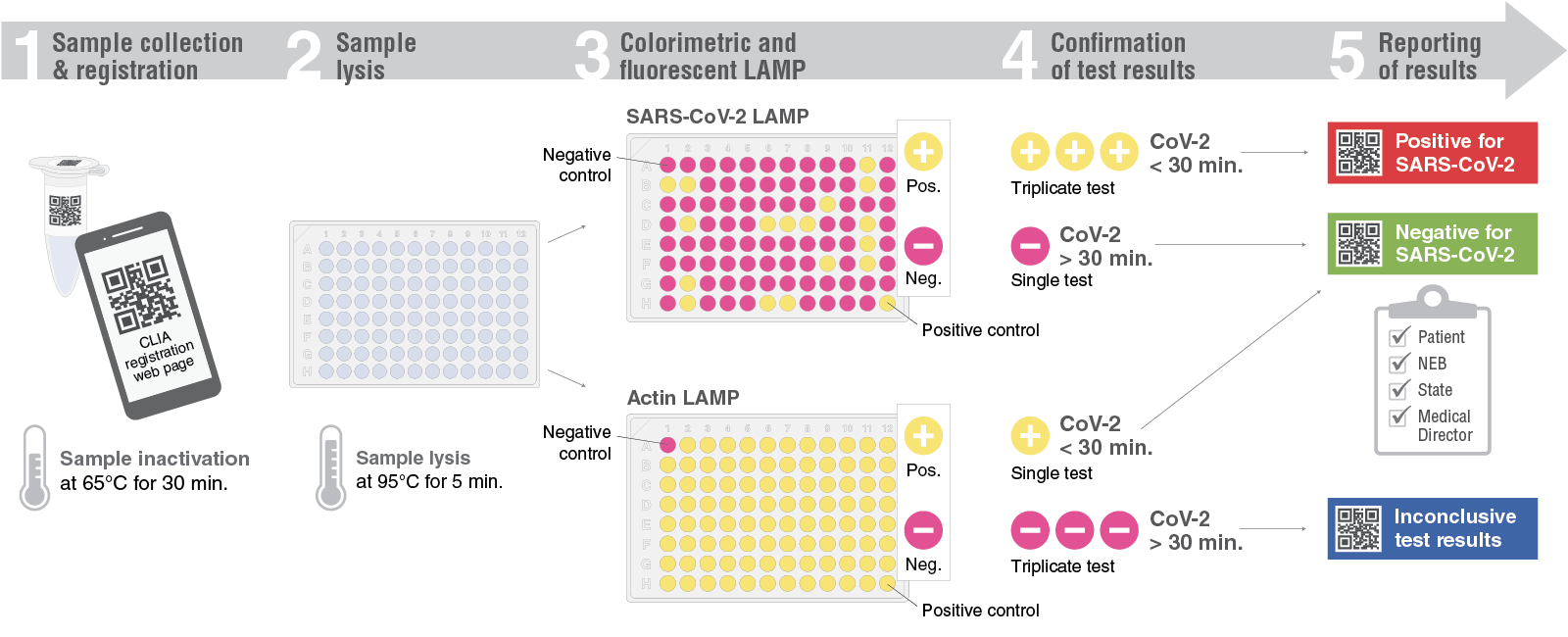
To ensure efficient sample processing with no risk to the NEB scientists conducting the tests, racks of samples are directly transferred to a 65°C incubator in our BSL2 lab for inactivation. Only after that step are samples handled, with equal volume saliva pipetted into a plate pre-filled with simple, colorimetric LAMP compatible lysis buffer inspired by similar efforts (9,11). Heating samples to 95°C in this buffer serves to release RNA from cells and viral particles, as well as inactivate the accompanying nucleases that would degrade the RNA. The same QR code identifier used to log the sample is also used in the lab to record its position on the plate with a photo of the tube rack. Samples are now ready for RT-LAMP: 2 μl of each sample is loaded into a SARS-CoV-2 LAMP plate and an Actin LAMP control plate.
For a streamlined testing workflow, we turned to standard 96-well qPCR instruments (Bio-Rad® CFX96™ Touch), which are used daily by various groups here at NEB, with two of these instruments dedicated specifically to the CLIA lab. The workflow for our SARS-CoV-2 Rapid Colorimetric LAMP Assay Kit involves processing 47 samples in one 96-well plate, with each sample tested in side-by-side reactions using either SARS-CoV-2 primers or internal control (human Actin) primers to confirm the presence of sample nucleic acid. Additionally, each plate includes one positive and one negative control. We adjusted this strategy slightly for higher throughput using the two designated instruments, with 94 samples distributed between Actin control and SARS-CoV-2 test plates, with positive and negative controls on each. Each plate is scanned using a simple flatbed document scanner before amplification to record the color of reactions, which all should begin as a bright pink color using our colorimetric LAMP mix. This is a built-in indicator of sample quality. In rare cases (<0.1% of samples) we observe an orange or yellow color after lysed saliva is added to the colorimetric LAMP mix, a sign of low pH, that may affect amplification performance. For detection of the LAMP reactions, we supplement the colorimetric mix with our LAMP Fluorescent Dye (NEB #B1700; equivalent to SYTO™-9). This is not required for measurement of LAMP because the resulting color change could easily be detected by eye or quantified using absorbance measurements, as with Color Health’s test (1), but we took advantage of existing in-house LIMS software built to take results from Bio-Rad CFX files for streamlined processing and analysis. We used amplification time thresholds to call reactions as either positive (detected actin, detected SARS-CoV-2), negative (detected actin, not detected SARS-CoV-2), or inconclusive (not detected actin). When the data from a plate is released, a clinical report is automatically generated with the date and important information for using the test result. Each user with a logged sample receives a report of their result via email.
Testing program results
The turnaround time for one complete run of 94 samples is ~2 hours, including the sample inactivation and RT-LAMP amplification steps. On average, the user experiences a longer wait time, as sample submission is of course variable (the first tube on the rack will wait a little while for the rack to fill up). We also collect samples from our multiple Massachusetts locations, and all samples received before a designated cutoff time on testing day are reported back to the employee the same day. Any initially positive or inconclusive sample is repeated in triplicate, and only if confirmed is reported to the user to ensure accurate results.
To date (March 1, 2022) we have processed 35,461 tests from 756 individuals, with 0.49% returned positive. In 0.11% of tests, we returned an inconclusive result, and employees who receive repeated inconclusive results generally have difficulty producing sufficient saliva. These employees are advised to either change the time of day for collection or adjust the process (e.g., drinking water immediately before collection), and these remedies have resulted in a high success rate. With our ability to rapidly test and release results, positive cases can be identified and kept off premises, and any close contacts are re-tested frequently. Follow-up re-testing is performed when samples are easily attained and we can observe the dynamics of infection over time. Negative test results are required before an on-site return (typically by ~Day 8). Trends in our results have matched the general population, with very few positives observed through summer 2021, and then increasing numbers in September-December, following the Delta wave, and increasing significantly with Omicron in December 2021-January 2022. These trends allow us to adjust our workplace policies and non-pharmaceutical interventions as needed to ensure the safest possible work environment, restricting the use of meeting rooms and requiring masks in response to case numbers onsite.
With the world-class sequencing expertise and resources at NEB, we additionally can perform next generation sequencing on RNA extracted from ever positive sample, validating the performance of our sequencing products with real-world variants and sublineages, and contributing to the global scientific knowledge base of SARS-CoV-2 spread. Following with the case trends, all samples detected and sequenced in Fall 2021 were determined to be B.1.617.2 (Delta), which completely transitioned mid-December to B.1.1.529 (Omicron).

Summary and outlook
With our CLIA authorization, established workflow and equipment for testing, and general appreciation for the additional layer of workplace safety that our testing provides, we plan to continue our SARSCoV-2 operations for the foreseeable future. And while our process works here at NEB, we continually advise others on the robustness and flexibility of RT-LAMP to accommodate other testing plans at a wide range of scales and settings. Examples include a similar workflow at Rice University (12), screening healthcare workers in Nicaragua (13), and an effort across multiple African countries spearheaded by ICGEB (14). Platforms as simple as heat and visual detection can be used for lowthroughput settings (15), small portable instruments for fieldable moderate throughput needs (16), and higher complexity workflows for high-throughput testing programs (17-18). With the experience and testing infrastructure we have built, setting up a test for potential new viral threats will be achievable with rapid timelines, and we will consider influenza or other target tests as needs arise. The ability to provide routine diagnostic testing has helped us safely continue our research and production operations through the ongoing pandemic, and we feel more environments and congregate settings adopting initiatives like ours would significantly benefit public health and viral control efforts.
View a PDF of this feature article
![]() For more information, read our preprint: ‘Development and Implementation of a Simple and Rapid Extraction-Free Saliva SARSCoV-2 RT-LAMP Workflow for Workplace Surveillance’ at https://www.medrxiv.org/content/10.1101/2022.03.11.22272282v1
For more information, read our preprint: ‘Development and Implementation of a Simple and Rapid Extraction-Free Saliva SARSCoV-2 RT-LAMP Workflow for Workplace Surveillance’ at https://www.medrxiv.org/content/10.1101/2022.03.11.22272282v1
References
- Emergency Use Authorization (EUA) Summary for the Color SARS-CoV-2 RT-LAMP Diagnostic Assay (https://www.fda.gov/media/138249/download).
- Notomi, T. et al. (2000) Nucleic Acids Res. 28, 12, PMID: 10871386.
- Poole, C.B. et al. (2012) PLoS Negl Trop Dis, 6, 12, e1948.
- Alhassan, A. et al. (2016) Mol Biochem Parasitol, 210, 1-2,10-12.
- Poole, C.B. et al. (2017) PLoS One, 12, 2, e016901.
- Zhang, Y. et al. (2020) (https://www.medrxiv.org/content/10.1101/2020.02.26.20028373v1.full.pdf+html).
- Zhang, Y et al. (2020) Biotechniques, 69, 3, 178-185.
- Vogels, C.B.F. et al. (2021) Med (NY) 2, 3, 263-280.
- Yan, Q. et al. (2021) Elife 10, e65113.
- Anahtar, M.N. et al. (2020) Open Forum Infect Dis, 8, 2, ofaa631.
- Rabe, B.A. and Cepko, C. (2020) Proc Natl Acad Sci USA, 117, 39, 24450-24458.
- Kundrod, K.A. et al. (2022) PLoS One, 17, 2 e0264130.
- Huete-Pérez, J. et al. (2021) PLoS One, 16, 1, e0246084.
- Baba, M.M. et al. (2021) EClinicalMedicine, 40, 101101.
- Lin, E.E. et al. (2022) PLoS One 17, 1, e0259886.
- Diaz, L.M. et al. (2021) J Biomol Tech, 32, 3, 158-171.
- Lalli, M.A. et al. (2021) Clin Chem, 67, 2, 415-424.
- Dao Thi, V.L. et al. (2020) Science Translational Medicine, 12, 556, eabc7075.

